goal
use genetic algorithm to find the local maxima for the function -
sin(x) - 0.2 * |x|defining problem
genotype
since the algorithm first initializes a population of chromosomes, we need to define what each chromosome will look like (it’s features/characteristics). in our case, each chromosome will have a single number x at random (in range 0-10) but our framework needs an enumerable type (basically a list) so we create a list of one single gene
def genotype do
gene = :rand.uniform(10)
%Chromosome{genes: [gene], size: 1}
endfitness function
next we define a fitness function which returns fitness of a chromosome, which acts as a heuristic or evaluation criteria. for that we apply the function we are trying to find the maxima for to the gene of the chromosome and return the result
def fitness_function(chromosome) do
x = hd(chromosome.genes) # getting top element from the list
Math.sin(x) - 0.2 * abs(x)
endtermination function
termination function signals the end of the algorithm. in our case it ends when number of generations hit 10, cause that’s when convergence in population is mostly observed (can be run longer but results won’t change much)
def terminate?(population, generation) do
plot_generation(population, generation)
generation == 10
endgraph plotting (optional)
graph plotting uses gnuplot to create graphs using gnuplot-elixir library (kindof like how we use matplotlib in python) to observer how each generation of chromosomes performs on the graph
defp plot_generation(population, generation) do
curve =
Enum.map(-100..100, fn x ->
x = x / 10.0
[x, :math.sin(x) - 0.2 * abs(x)]
end)
points =
Enum.map(population, fn c ->
[hd(c.genes), c.fitness]
end)
best = hd(population)
best_point = [[hd(best.genes), best.fitness]]
{:ok, _} =
Gnuplot.plot(
[
[:set, :title, "Generation #{generation}"],
Gnuplot.plots([
["-", :with, :lines, :lt, 2, :title, "sin(x) - 0.2|x|"],
["-", :with, :points, :pt, 7, :ps, 1.5, :lc, "orange", :title, "Population"],
["-", :with, :points, :pt, 5, :ps, 2, :lc, "green", :title, "Best"]
])
],
[curve, points, best_point]
)
endgraphs - plots from generation 1 to generation 10 where entire population converges at a single point
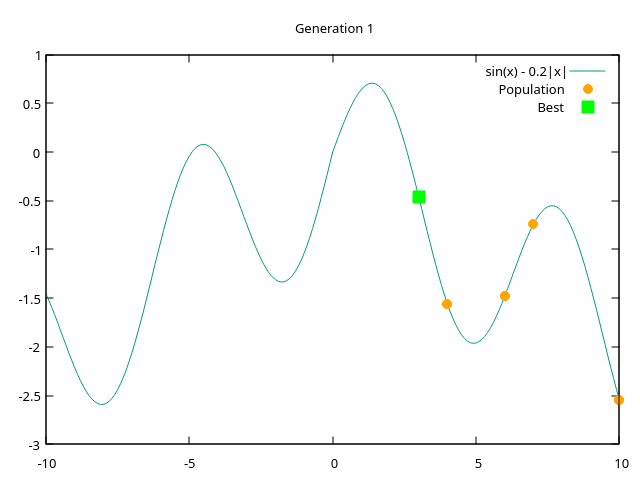 | 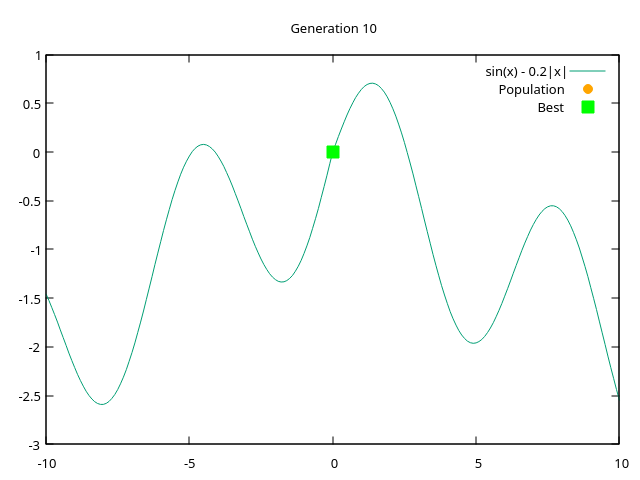 |
|---|
working of the framework
now coming to what is happening behind the scenes in the framework,
initialization
first step in any genetic algorithm is to initialize the population. so here we just create a population of size 10 (can be larger as well) where each chromosome has the genotype we defined earlier
def initialize(genotype, opts \\ []) do
population_size = Keyword.get(opts, :population_size, 100)
population = Enum.map(1..population_size, fn _ -> genotype.() end)
population
endevaluation
next comes evaluating each chromosome and sorting them based on their fitness/heuristic/evaluation criteria
def evaluate(population, fitness_function, _opts \\ []) do
population
|> pmap(fn chromosome ->
fitness = fitness_function.(chromosome)
age = 1
%Chromosome{genes: chromosome.genes, fitness: fitness, age: age}
end)
|> Enum.sort_by(& &1.fitness, :desc)
endselection
now that we have a list of chromosomes, we select parents for crossover. these parents that will be combined to create new solutions the goal of selection is to pick some parents that can easily be combined to create better solutions
def select(population, opts \\ []) do
select_rate = Keyword.get(opts, :selection_rate, 0.8)
n = round(length(population) * select_rate)
n = if rem(n, 2) == 0, do: n, else: n - 1
parents = tournament(population, n)
leftover =
population
|> MapSet.new()
|> MapSet.difference(MapSet.new(parents))
parents =
parents
|> Enum.chunk_every(2)
|> pmap(&List.to_tuple(&1))
{parents, MapSet.to_list(leftover)}
end
def tournament(population, n) do
tournsize = :rand.uniform(n)
0..(n - 1)
|> Enum.map(fn _ ->
population
|> Enum.take_random(tournsize)
|> Enum.max_by(& &1.fitness)
end)
endhere we are using a selection technique called tournament selection, which works kinda like this -
- choose a pool of n chromosomes where n is the tournament size
- choose the fittest chromosome from the tournament
- repeat
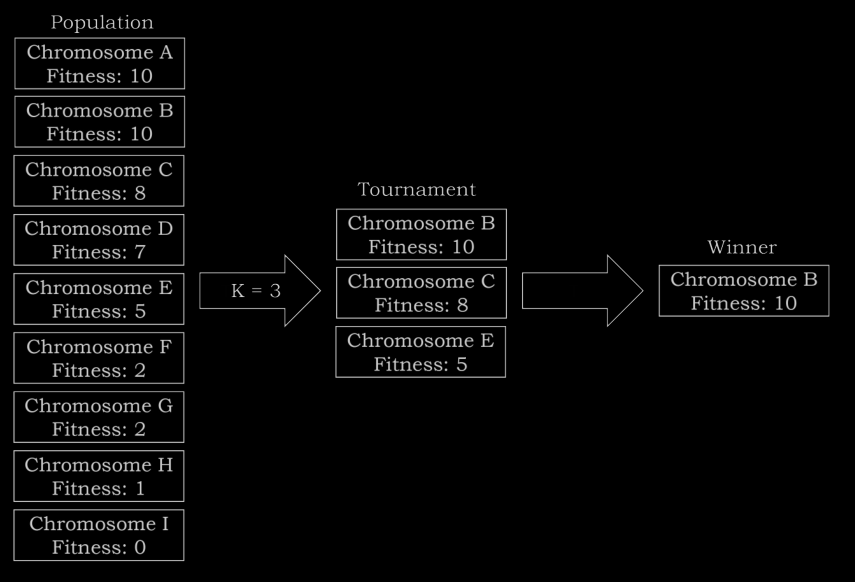
the selection rate can be set depending on the requirement but the leftovers created are reinserted as to preserve the population size (may also increase or decrease depending on how mutation rate is configured)
crossover
now to generate new solution we take two or more parent chromosomes and produce two or more child chromosomes. here we are using blend crossover. it linearly mixes the parent genes using a random shift factor (scaled by alpha) to generate two offspring as weighted combinations, then clamps the results within the specified bounds (in our case between -10 to 10)
def crossover(population, opts \\ []) do
population
|> Enum.reduce(
[],
fn {p1, p2}, acc ->
{c1, c2} = apply(crossover_fn, [p1, p2, opts])
Utilities.Genealogy.add_chromosome(p1, p2, c1)
Utilities.Genealogy.add_chromosome(p1, p2, c2)
[c1, c2 | acc]
end
)
end
def blend(p1, p2, opts \\ []) do
p1_1 = Enum.at(p1.genes, 0)
p2_1 = Enum.at(p2.genes, 0)
alpha = Keyword.get(opts, :alpha, 1)
{min, max} = {Keyword.get(opts, :min, -10), Keyword.get(opts, :max, 10)}
shift = (0.1 + 0.2 * alpha) * :rand.uniform() - alpha
c1 = (0.1 - shift) * p1_1 + shift * p2_1
c2 = shift * p1_1 + (0.1 - shift) * p2_1
{
%Chromosome{genes: [constraints(c1, min, max)], size: 1, age: 0},
%Chromosome{genes: [constraints(c2, min, max)], size: 1, age: 0}
}
end
defp constraints(p, min, max) do
p = if p > max, do: max, else: p
p = if p < min, do: min, else: p
p
endmutation
since there is a possibility of early convergence in population we randomly mutate certain number of chromosomes to introduce randomness in the population. here we select a random subset of the population (based on mutation rate) and replace each selected chromosome’s genes with values sampled from a gaussian distribution defined by its own mean and variance
def mutation(population, opts \\ []) do
mutation_rate = Keyword.get(opts, :mutation_rate, 0.05)
n = floor(length(population) * mutation_rate)
population
|> Enum.take_random(n)
|> Enum.map(&gaussian(&1, opts))
end
def gaussian(chromosome, _opts \\ []) do
mu = Enum.sum(chromosome.genes) / length(chromosome.genes)
sigma =
chromosome.genes
|> Enum.map(fn x -> (mu - x) * (mu - x) end)
|> Enum.sum()
|> Kernel./(length(chromosome.genes))
genes =
chromosome.genes
|> Enum.map(fn _ -> :rand.normal(mu, sigma) end)
%Chromosome{genes: genes, size: chromosome.size}
endgenerating the solution
finally we run the algorithm and the graphs mentioned in [termination function] get generated as each generation passes and the algorithm ends at generation 10
(configurations for the algorithm)
{_soln, _gen} =
Genetic.run(
Sin, # module name of thje problem
population_size: 10,
selection_rate: 0.8,
mutation_rate: 0.1,
# configurations for crossover
alpha: 1,
min: -10,
max: 10,
)conclusion
well, that was in short how a problem can be solved or rather a desired outcome can be found using genetic algorithm and how it sort of all comes together in my framework. thx for reading!
(unrelated but how the graph would look if it was 3d (beautiful right?))
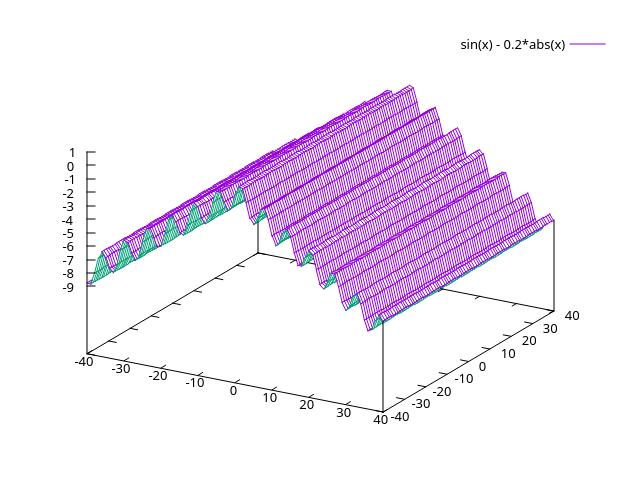
comments